The showyourwork.yml config file
Contents
The showyourwork.yml config file#
This is the configuration file for showyourwork!, allowing you to customize several aspects of the workflow. Below is a list of all available options.
datasets#
Type: mapping
Description: A mapping declaring static datasets to be downloaded from Zenodo or Zenodo Sandbox. Nested under this keyword should be a sequence of mappings labeled by the deposit version DOIs of Zenodo or Zenodo Sandbox datasets. See below for details.
Required: no
Example:
The following block shows how to tell showyourwork! about two files,
TOI640b.json and KeplerRot-LAMOST.csv, each of which is hosted
on a Zenodo deposit with a different version DOI. Note that the user should
separately provide dependencies information for each of these
files, so showyourwork! knows which scripts require these files.
datasets:
10.5281/zenodo.6468327:
contents:
TOI640b.json: src/data/TOI640b.json
10.5281/zenodo.5794178:
contents:
KeplerRot-LAMOST.csv: src/data/KeplerRot-LAMOST.csv
See below for the syntax of the contents section of the datasets mapping.
datasets.<doi>#
Type: mapping
Description: The Zenodo or Zenodo Sandbox version DOI for the deposit.
Note
Zenodo makes a distinction between version DOIs and concept DOIs. Version DOIs are static, and tied to a specific version of a deposit (the way you’d expect a DOI to behave); this is what you should provide here. Concept DOIs, on the other hand, point to all versions of a given record, and always resolve to the latest version. Check out the sidebar on the web page for this sample deposit:
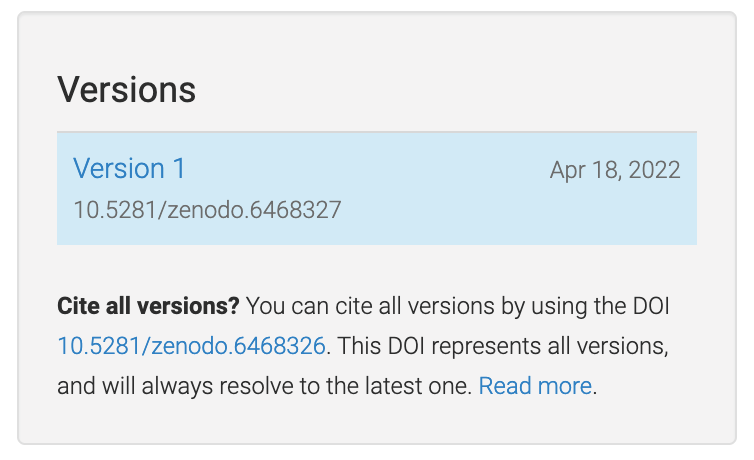
You can see that the DOI 10.5281/zenodo.6468327 corresponds to a specific version (1)
of the deposit, while the DOI 10.5281/zenodo.6468326 corresponds to all versions of
the deposit (it’s listed under “Cite all versions?”).
The former is a “version” DOI, while the latter is a “concept” DOI.
You can read more about that in the Zenodo docs.
Required: no
Example:
If the version DOI for a deposit containing the file TOI640b.json is 10.5281/zenodo.6468327,
we would specify the following in the config file:
datasets:
10.5281/zenodo.6468327:
contents:
TOI640b.json: src/data/TOI640b.json
See below for the syntax of the contents section of the datasets mapping.
datasets.<doi>.contents#
Type: mapping
Description: Specifies a mapping between files in a Zenodo or Zenodo Sandbox deposit and local
files. The contents field must contain key-value pairs of the form
remote-file: path-to-local-file
where remote-file is the name of the file on the remote (the Zenodo deposit)
and path-to-local-file is the path to the file on disk, relative to the
top level of the repository. The path-to-local-file may be omitted, in which
case the file name is preserved and the file is downloaded to the default
destination (see the option of the same name below).
If the remote file is a zipfile or a tarball, instead of a local path, users may provide a directory tree mapping that specifies the contents of the tarball and where they should be extracted to. The workflow will automatically extract them. See the example below for details.
Note
The contents section need only specify files used by the workflow; if
there are additional files in the Zenodo deposit that are not needed by
the workflow, they need not be listed. However, files that required by
the workflow must be listed explicitly; glob syntax is not allowed.
Required: no
Example:
The following example shows all the various ways in which Zenodo files can be downloaded, extracted, and mapped to local files:
datasets:
10.5281/zenodo.6468327:
destination: src/data/TOI640 # default folder to extract files to
contents:
README.md: # auto extracted to `src/data/TOI640/README.md`
TOI640b.json: src/data/TOI640/planet.json # rename the extracted file, just for fun
images.tar.gz: # remote tarballs behave like folders w/ same name
README.md: # auto extracted to `src/data/TOI640/images/README.md`
S06: # subfolder
image.json: src/data/TOI640/S06.json # rename and change destination folder
S07: # subfolder
image.json: src/data/TOI640/S07.json # rename and change destination folder
lightcurves.tar.gz: # another tarball
lightcurves: # files are nested inside `lightcurves` in this tarball
README.md: # auto extracted to `src/data/TOI640/lightcurves/lightcurves/README.md`
S06: # subfolder
lc.txt: src/data/TOI640/S06.txt # rename and change destination folder
S07: # subfolder
lc.txt: src/data/TOI640/S07.txt # rename and change destination folder
Recall that users must separately provide dependency information for each of these files via the dependencies key.
datasets.<doi>.destination#
Type: str
Description: The default destination to extract the contents of the Zenodo deposit to.
Required: no
Default: src/data
Example:
The following will extract all files in the Zenodo deposit with doi 10.5281/zenodo.6468327
to src/data (subfolders will be preserved).
datasets:
10.5281/zenodo.6468327:
destination: src/data
dependencies#
Type: list
Description: List of dependencies for each script. Each entry should be
the path to a script (either a figure script or the TeX manuscript itself)
relative to the repository root. Following each entry, provide a list of
all files on which the script depends. These dependencies may either be
static (such as helper scripts) or programmatically generated (such as
datsets downloaded from Zenodo). In the latter case, instructions on how
to generate them must be provided elsewhere (either via the zenodo key
below or via a custom rule in the Snakefile). In both cases, changes
to the dependency will result in a re-run of the section of the workflow that
executes the script.
Required: no
Default: []
Example:
Tell showyourwork! that the figure script my_figure.py depends on
the helper script utils/helper_script.py:
dependencies:
src/scripts/my_figure.py:
- src/scripts/utils/helper_script.py
You can also specify a dependency on a programmatically-generated file:
dependencies:
src/scripts/fibonacci.py:
- src/data/fibonacci.dat
provided data/fibonacci.dat is defined in a zenodo deposit (see below)
or instructions for generating it are provided in the Snakefile.
Finally, dependencies of the manuscript file are also allowed:
dependencies:
src/ms.tex:
- src/answer.tex
ms#
Type: str
Description: Path to the main TeX manuscript. Change this if you’d prefer to
name your manuscript something other than src/tex/ms.tex. Note that you should still
keep it in the src/tex directory. Note also that the compiled PDF file will
have the same name (e.g., src/tex/article.tex will generate article.pdf
in the repository root) .
Required: no
Default: src/tex/ms.tex
Example:
ms: src/tex/article.tex
overleaf#
Type: mapping
Description: Settings pertaining to Overleaf integration. See below for details, and make sure to check out Overleaf integration.
Required: no
Example:
overleaf:
id: 62150dd16134ef045f81d1c8
auto-commit: true
push:
- src/tex/figures
pull:
- src/tex/ms.tex
- src/tex/bib.bib
overleaf.id#
Type: str
Description: The id of the Overleaf project to integrate with. This can be obtained from the URL of the project, e.g.:
https://www.overleaf.com/project/6262c032aae5421d6d945acf
in this case, the id is 6262c032aae5421d6d945acf.
Warning
Please read the Overlaf integration docs before manually adding/changing this value, as you could risk losing changes to your local document or to your Overleaf document the next time you build!
Required: no
Example:
overleaf:
id: 62150dd16134ef045f81d1c8
overleaf.pull#
Type: bool
Description: A list of files and/or folders to be pulled from the Overleaf project before every build. These should be files that are only ever modified on Overleaf, such as the TeX manuscript and other TeX files. Paths should be relative to the top level of the repository. Exact names are required; no glob syntax allowed.
Required: no
Default: []
Example:
overleaf:
pull:
- src/tex/ms.tex
- src/tex/bib.bib
overleaf.push#
Type: bool
Description: A list of files and/or folders to be pushed to the Overleaf project after every build. These should be files that are programmatically generated by the build, such as the figure files. Paths should be relative to the top level of the repository. Exact names are required; no glob syntax allowed.
Required: no
Default: []
Example:
overleaf:
push:
- src/tex/figures
require_inputs#
Type: bool
Description: If there is no valid rule to generate a given output file
(because of, e.g., a missing input file), but the output file itself is present on disk,
Snakemake will not by default raise an error. This can be useful for running
workflows locally, but it can compromise the reproducibility of a workflow when
a third party attempts to run it. Therefore, the default behavior in showyourwork!
is to require all output files to be programmatically generatable when running
the workflow, even if the output files exist on disk already. Otherwise, an
error is thrown. Set this option to false to override this behavior.
Required: no
Default: true
Example:
require_inputs: true
run_cache_rules_on_ci#
Type: bool
Description: Allow cacheable rules to run on GitHub Actions if the cached
output is not available? Default is false, which prevents potentially
computationally expensive rules from running on the cloud. In this case,
cache misses result in an error when running on GitHub Actions only.
Required: no
Default: false
Example:
run_cache_rules_on_ci: false
scripts#
Type: mapping
Description: Mapping of script extensions to instructions on how to execute
them to generate output. By default, showyourwork! expects output files
(e.g., figures or datasets) to
be generated by executing the corresponding scripts with python. You can add custom
rules here to produce output from scripts with other extensions, or change
the behavior for executing python scripts (such as adding command line
options, for instance). Each entry under scripts should be a file extension,
and under each one should be a string specifying how to generate the output file
from the input script. The following placeholders are recognized by showyourwork!
and expand as follows at runtime:
{script}: The full path to the input script.{output}: The full path to the output file (i.e., the generated figure). If the script generates more than one file, this expands to a space-separated list of outputs.{datasets}: A space-separated list of all the Zenodo datasets required by the current script.{dependencies}: A space-separated list of all the dependencies (including datasets) of the current script.
Required: no
Default: The default behavior for python scripts corresponds to the
following specification in the yaml file:
scripts:
py:
python {script}
That is, python is used to execute all scripts that end in .py.
Example: We can tell showyourwork! how to generate figures by executing a Jupyter notebook as follows:
scripts:
ipynb:
jupyter execute {script}
style#
Type: mapping
Description: Specifies custom modifications to the article stylesheet.
Required: no
style.show_git_sha_or_tag#
Type: bool
Description: Show the git SHA in the article PDF header. If the HEAD commit corresponds to a git tag, show the tag name in the header.
Required: no
Default: false
Example:
style:
show_git_sha_or_tag: true
verbose#
Type: bool
Description: Enable verbose output? Useful for debugging runs. By default,
showyourwork! suppresses nearly all Snakemake output, sending it directly
to the log file (see Logging). Setting verbose: true results in all
Snakemake output being printed to the screen as well. Note that you can
crank up the verbosity even more by passing the --verbose argument to
snakemake build, which makes Snakemake itself more talkative.
Required: no
Default: false
Example:
verbose: true
version#
Type: str
Description: The version of the showyourwork! package required to build
the article, populated automatically when showyouwork setup is run. Users
may, however, change this to upgrade/downgrade to a different version of the
package. Options are:
any pip-installable version number (e.g.,
0.3.0)a 5-character (short) or 40-character (long) GitHub commit SHA (e.g,
abcde) corresponding to a specific commit to the github.com/showyourwork/showyourwork repo
Required: yes
Example:
version: 0.3.0
