Introduction¶
The showyourwork workflow is intended to help authors publish the code that generated
the figures and results in a scientific article. It ensures that the compiled article
PDF is always in sync with all of the code used to generate it. It does this
automatically—and seamlessly—with the help of the Snakemake workflow management system,
the tectonic typesetting engine, and
Github Actions CI.
The showyourwork philosophy
Scientific papers should exist as GitHub repositories comprised of LaTeX files, figure scripts, rules to access datasets, a platform/environment specification, and nothing else. Anyone should be able to re-generate the article PDF from scratch at the click of a button.
Within the showyourwork workflow, scientific articles exist as GitHub repositories
with a specific layout. Whenever new commits are pushed to the remote
repository, a GitHub action is triggered that automatically builds the article from the
input figure scripts, manuscript files, and
conda environment file,
following the instructions specified in the Snakefile:
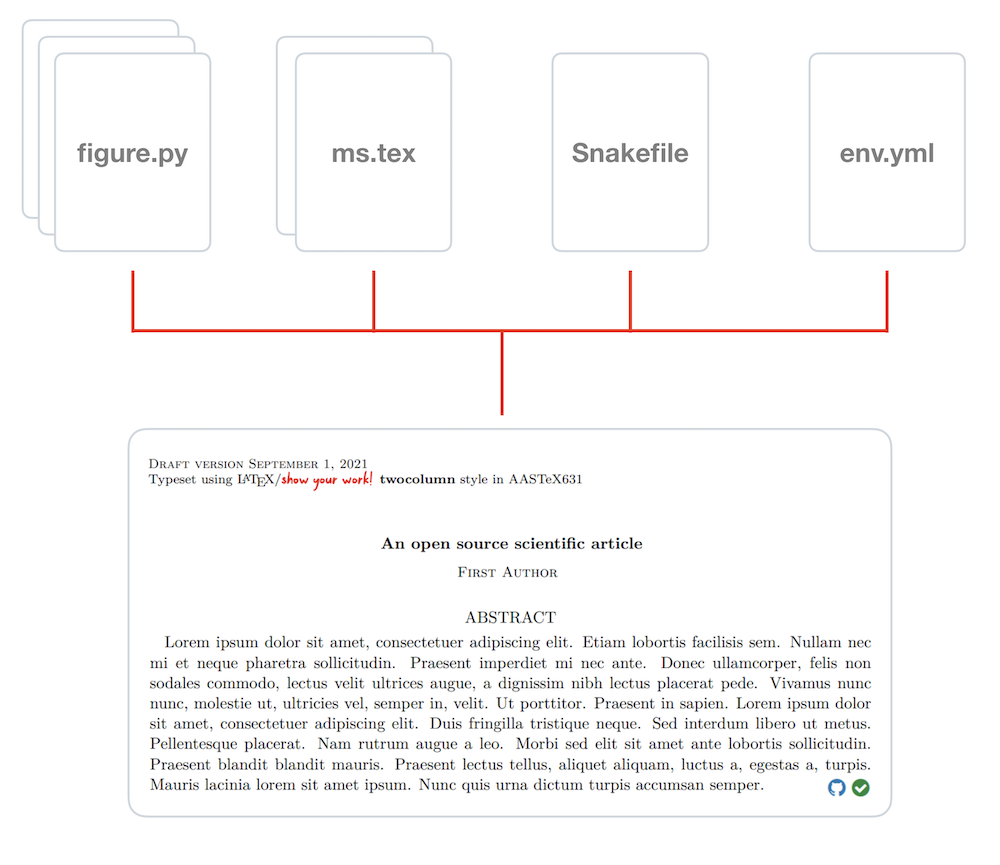
The article PDF (along with a tarball containing all of the output) is then pushed to a special branch
(usually main-pdf) on the repository. This article is decorated with badges linking to the exact
versions of the files on GitHub used to generate it.
Thanks to the magic of Snakemake, showyourwork is both lightweight—it should work out-of-the-box for most users—and highly
customizable. It also uses intelligent caching to never re-run things it doesn’t have to (like figure scripts that haven’t changed),
even when running on the cloud.
To get started with showyourwork, check out the quickstart tutorial.
You should also read about
the showyourwork GitHub action, how to build your article locally,
and how to customize your workflow.
You should also spend some time browsing through the FAQs page. Since showyourwork
is itself a work in progress, new features are still being added frequently. If you spot a bug,
have a question, or would like showyourwork to do something it doesn’t currently support,
please feel free to raise a GitHub issue.
